Biohackathon fixes#2695
Conversation
|
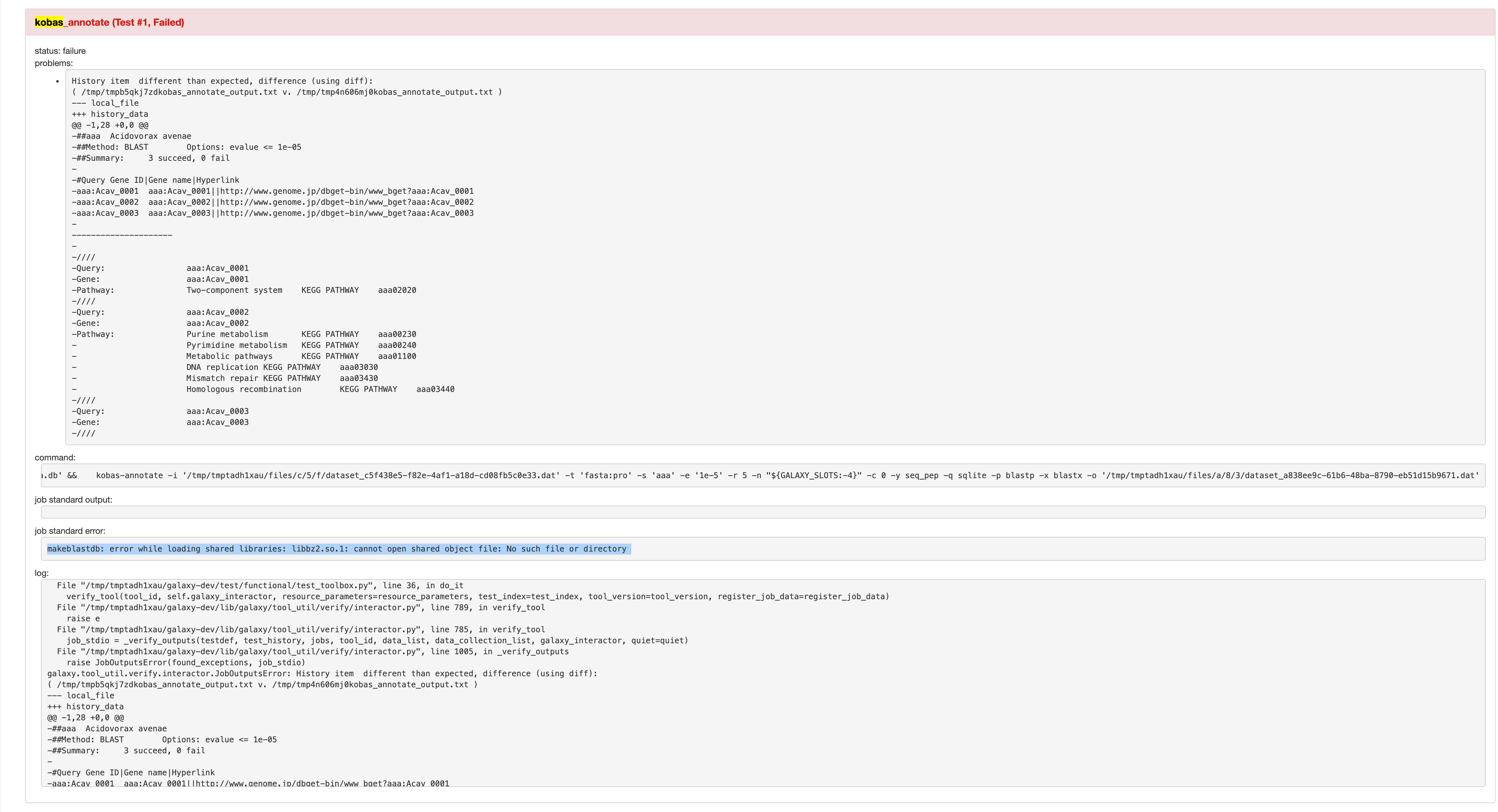
Not sure why kobas_annotate test does not produce output for 20', ping @blankclemens . |
|
Anyway, probably better to do smaller PRs here to not get slowed down |
|
I've replicated the After more than 20' it still hasn't solved the environment. |
|
@nsoranzo can we bump the conda recipe and get a new build? |
|
Yeah, it's been doing that all night on my laptop (I just added bzip2). The 2.1.1 recipe is not in the branch anymore, should we add that back ? |
|
@mvdbeek I'm going to push a fix in a bit, adding r-base 3.4 as additional requirement for kobas tools. |
|
Cool! If you want you could rebase out the mothur changes so that we actually get a chance at getting the tool tests to finish on time. |
With the wrong profile tag dependencies wouldn't get resolved.
Uses regex for match rna_star data manager output. Uses yield to add star requirements, as macro is re-used in star tool.
The test comes from a time when we didn't test data managers yet,
so I think this probably didn't work. fwiw the output looks correct:
```
{"data_tables": {"all_fasta": [{"dbkey": "refseq_plastid.97.protein", "name": "RefSeq plastid Release 97 protein (2018-03-14)", "path": "/tmp/tmp_uJxfm/job_working_directory/000/1/dataset_8405ba59-24ae-4600-b132-38b696e22e49_files/plastid.97.protein.fasta", "value": "refseq_plastid.97.protein"}]}}
```
Co-Authored-By: Nicola Soranzo <nicola.soranzo@gmail.com>
b6815ec to
2030b8a
Compare
Done! |
This PR collects all fixes done so far during the biohackathon and in preparation to test all tools on this repo.
FOR CONTRIBUTOR: